
Tcl
1 Jun 2022 - Using this module, the "Materials" parameter-bundle of a label-field is created and filled with an ID and name for each label-value. Learn more

Python (pyscro)
13 May 2022 - This Xtra automatically sets the scalebar parameters to comply with the requirements for scientific papers. Learn more
Tcl
13 May 2022 - The coordinate-axes displayed using the "Axes" module cannot be scaled easily. Using this Xtra, you can easily change the lengths of the axes and the location of the displayed origin. Learn more

High Content Screening Plate Manager
Python (pyscro), Tcl
13 May 2022 - Load, visualize, and process multi-channel fields from Thermo Scientific CellInsight CX5, CX7 LED, and CX7 LZR HCA instruments and HCS Studio software. Learn more

Bone Segmentation Workflow for Murine Hindpaw
Recipe ISP, IVP, A2D, Demo, Data
28 Apr 2022 - Semi-automatic workflow and recipe for high-throughput bone segmentation of murine hindpaw micro-CT datasets. Learn more
Python (pyscro)
28 Apr 2022 - This module exports an image of color-washed data. Learn more
How to Segment Cavities Using the Compute Ambient Occlusion Module
Demo
12 Apr 2022 - This tutorial demonstrates how to segment pores or cavities using the Compute Ambient Occlusion module. Learn more

How to Perform Trabecular and Cortical Bone Segmentation
Demo
12 Apr 2022 - This tutorial demonstrates how to use the Compute Ambient Occlusion module to perform the segmentation of trabecular and cortical bone, as well as of the pore space, and then how to use the Arithmetic module to aggregate those results in one label field. Learn more

XPand module
6 Apr 2022 - This module performs the thinning of a binary image slice-by-slice. Learn more

Python (pyscro), Project (hx, pgo)
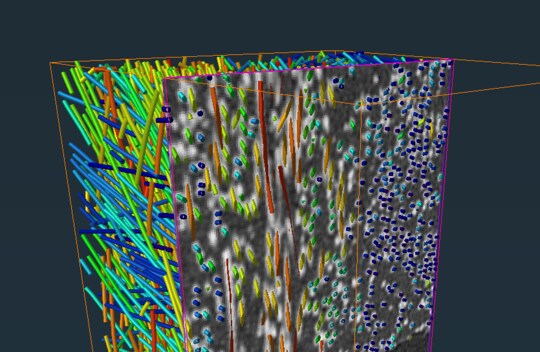
24 Feb 2022 - The Slab View display module allows you to display only a slab from any kind of 3D visualization. Learn more

How to Use the Surface View Module and Create Elaborate Surface Renderings
Demo
30 Dec 2021 - This short video demonstrates the basic use of the Surface View module, and how to combine several Surface View modules to create elaborate renderings. Learn more

Facies Classification from Well Logs
Python (pyscro), Demo
3 Dec 2021 - Facies (rock type) classification from well logs using supervised machine learning algorithms. Learn more

How to Customize Amira-Avizo with moduleExtender
Tcl, Misc
25 Nov 2021 - Are there module settings you always need to change during your daily work? This Xtra describes how you can permanently change default settings of modules in Amira-Avizo. Learn more

Tcl
18 Nov 2021 - Module Mask Map By Molecule lets you mask an EM map of a macro molecule by an atomic model of the entire or partial molecule. This can be useful when validating EM maps from SPA with published models from the Protein Data Bank. Learn more

kMeans Clustering on Label Analysis (Python)
Python (pyscro)
18 Nov 2021 - This Xtra implements kMeans clustering on a label analysis spreadsheet. Separated objects can be clustered based on any measure group that is inbuilt or user customized. Learn more

Differential Interference Contrast (DIC) Image Segmentation and Quantification using Deep Learning
Recipe (hxrecipe), Deep learning model, Demo
18 Nov 2021 - This workflow uses deep learning to segment images obtained by Differential Interference Contrast (DIC) microscopy. Nuclei and mitochondria are segmented and counted cell per cell. Learn more

Porosity Volume Fraction Analysis
Project (hx, pgo), Demo, Data
18 Nov 2021 - This tutorial demonstrates the steps to quantify porosity volume fraction and distribution. Learn more

Python (pyscro)
18 Nov 2021 - Add bookmarks to your project, including status and camera, so you can easily retrieve a given visualization. You can then navigate through the bookmarks. Learn more

Tcl, Demo
18 Nov 2021 - This module can be used to facilitate the alignment of datasets using pairs of landmarks. The video tutorial demonstrates this technique. Learn more

Inter Time Steps Recipe Player
Recipe (hxrecipe), Python (pyscro), Project (hx, pgo), Demo, Data
18 Nov 2021 - The Inter Time Steps Recipe Player module executes a two-input recipe to a specific pair of two successive time steps of a time series dataset, or to all its pairs of two successive time steps. Learn more

CMake Usage to Create XPand C++ API Modules
Demo
18 Nov 2021 - Starting with version 2021.2, the use of CMake is required to create solutions that implement the Amira Avizo 3D Pro extension modules with the XPand C ++ API. This video demonstrates what needs to be done to create a solution for MS Visual Studio. Learn more